Abstract
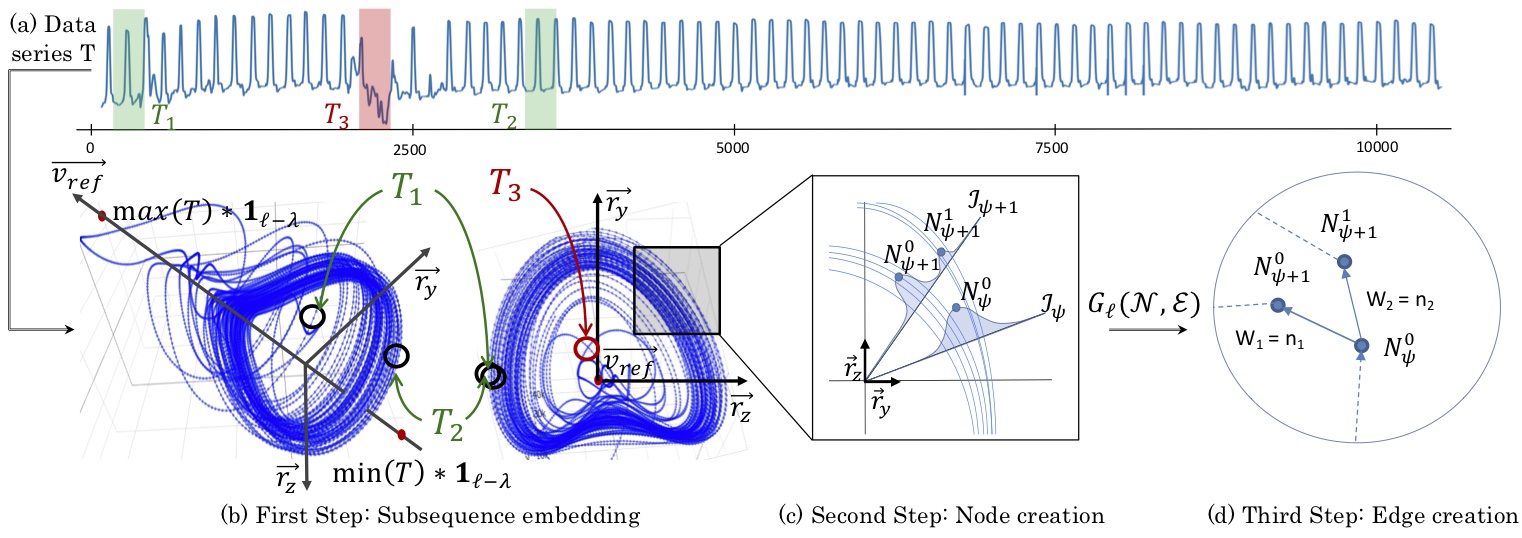
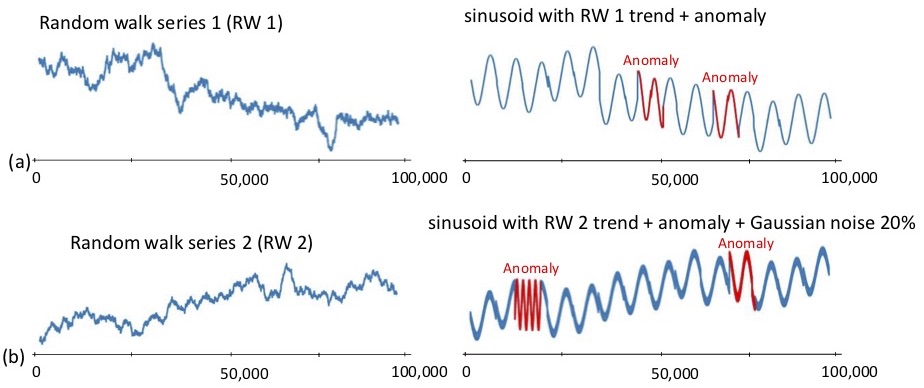
Subsequence anomaly detection in long sequences is an important problem with applications in a wide range of domains. However, the approaches that have been proposed so far in the literature have severe limitations: they either require prior domain knowledge that is used to design the anomaly discovery algorithms, or become cumbersome and expensive to use in situations with recurrent anomalies of the same type. In this work, we address these problems, and propose an unsupervised method suitable for domain agnostic subsequence anomaly detection. Our method, Series2Graph, is based on a graph representation of a novel low-dimensionality embedding of subsequences. Series2Graph needs neither labeled instances (like supervised techniques), nor anomaly-free data (like zero-positive learning techniques), and identifies anomalies of varying lengths. The experimental results, on the largest set of synthetic and real datasets used to date, demonstrate that the proposed approach correctly identifies single and recurrent anomalies without any prior knowledge of their characteristics, outperforming by a large margin several competing approaches in accuracy, while being up to orders of magnitude faster.
P. Boniol and T. Palpanas, Series2Graph: Graph-based Subsequence Anomaly Detection in Time Series, PVLDB (2020)
P. Boniol and T. Palpanas and M. Meftah and E. Remy, GraphAn: Graph-based Subsequence Anomaly Detection, demo PVLDB (2020)